The “Anakin” method: Using artificial intelligence to fight disease

A team of chemists have created new software — inspired by Star Wars — that will help researchers test and develop medicine faster.
Growing up in small-town Central Alabama, Justin Smith was drawn to Legos and science fiction books and films, like Star Wars and Star Trek — things that allowed him to imagine futuristic realities.
He’s still building works inspired by his love of science fiction, but they’re very real.
For the past year, the University of Florida doctoral student and colleagues have been developing software using artificial intelligence, specifically a neural network, that can be taught and even learn on its own. The developers hope the new software will lead to faster, cheaper and more accurate testing of treatments and potential cures for diseases and viruses, like diabetes and HIV.
“I wanted to make a difference in the way medicine is developed, ” said Smith, who studies at UF’s chemistry department and recently introduced the software in a study. “With this software, drug companies can identify a target protein, like those that drive the HIV virus, and test binding molecules to proteins that can stop the protein from working, essentially stopping the disease in its tracks.”
The software is called ANAKIN-ME (Accurate NeurAl networK engINe for Molecular Energies) or Ani for short. And describing Ani as “fast” is an understatement.
In order to bind a molecule to a protein, researchers need to understand the molecule’s energy. Smith taught the neural network to calculate energies and forces in molecules hundreds of thousands of times faster than the previous most accurate method — and much more accurately than the quickest methods that currently exist.
The combination of speed, accuracy and low cost will allow researchers to run experiments that were previously impossible, said Smith’s mentor Adrian Roitberg, a UF chemistry professor and study co-author.
“Drugs that used to take decades to test can be tested in months,” said Roitberg, who emphasized the software’s accuracy.
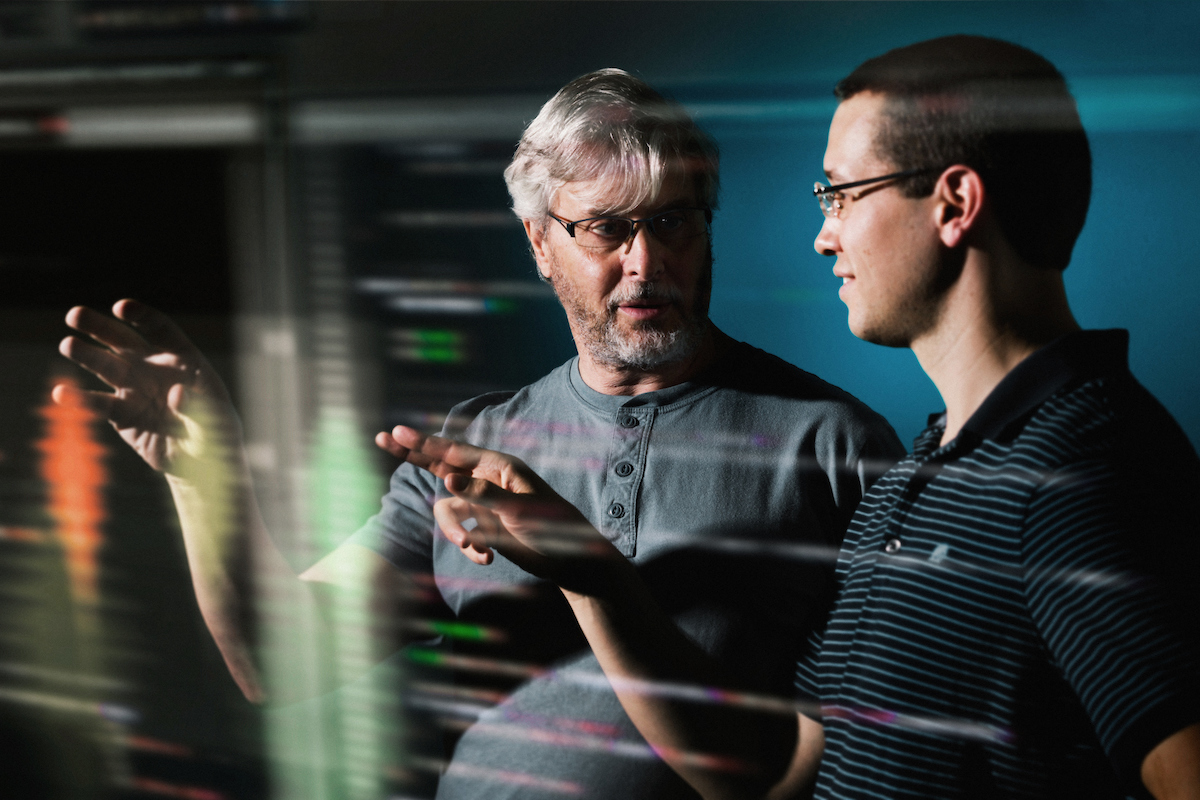

Justin Smith (right) and Adrian Roitberg discuss the new method.
The software might also help engineers and manufacturers develop materials like solar cells and plastics. The network is useful when building things, as it can identify the right materials and take away much of the trial and error, Roitberg said.
The problem with some previous methods is that they were not adapted to the way molecules move and bend, while others that were adapted to movement failed to work in complex chemical environments.
So, Smith took a different approach.
“The idea was to design a method that works the same way they teach cars to drive. You don’t teach a car to drive down a straight road and then expect it to take corners, for instance, “ said Smith, a UF Foundation graduate fellowship recipient. “Because molecules move, I realized we needed to design a way to teach the neural network that things are not rigid.”
The speed at which Ani computes might allow researchers to test huge numbers of possible molecule and protein connections, or study big, complex molecules.
Though it’s already advanced, Ani is still learning. It knows organic chemistry, but eventually the network might be able to train itself to perform even more accurately.
And as for the reason Ani’s creators named it after a Star Wars character, Roitberg said, “In those movies, computers were solving problems long before words like artificial intelligence or machine learning existed. So it’s a shout out to science fiction and the way it spurred interest in computers solving real-world problems.”
Olexandr Isayev with the University of North Carolina at Chapel Hill co-authored the study.